GREAT FLUIDIC SOLUTIONS
Competent products with expert advice
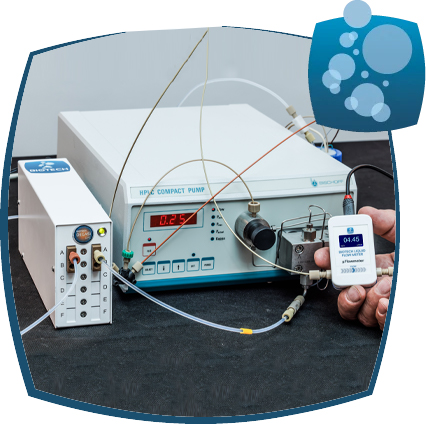
Biotech Fluidic provides a carefully selected range of products to design and improve high-precision systems for liquid analysis and processing at many different flow rates and solvent conditions. Among the trusted brands we offer are IDEX Health & Science, Systec, Upchurch, Rheodyne, Isolation Technologies, Mott Corporation, and Runge, plus a rapidly growing product portfolio originating from Biotech Fluidics. Together these products cover a wide range of requirements in liquid degassing, inline monitoring, separation tools, fluid management, and flow components.